![PDF] Easy quantitative assessment of genome editing by sequence trace decomposition | Semantic Scholar PDF] Easy quantitative assessment of genome editing by sequence trace decomposition | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/055277c5b235228474242c3562c4cad2e02627a7/4-Figure1-1.png)
PDF] Easy quantitative assessment of genome editing by sequence trace decomposition | Semantic Scholar

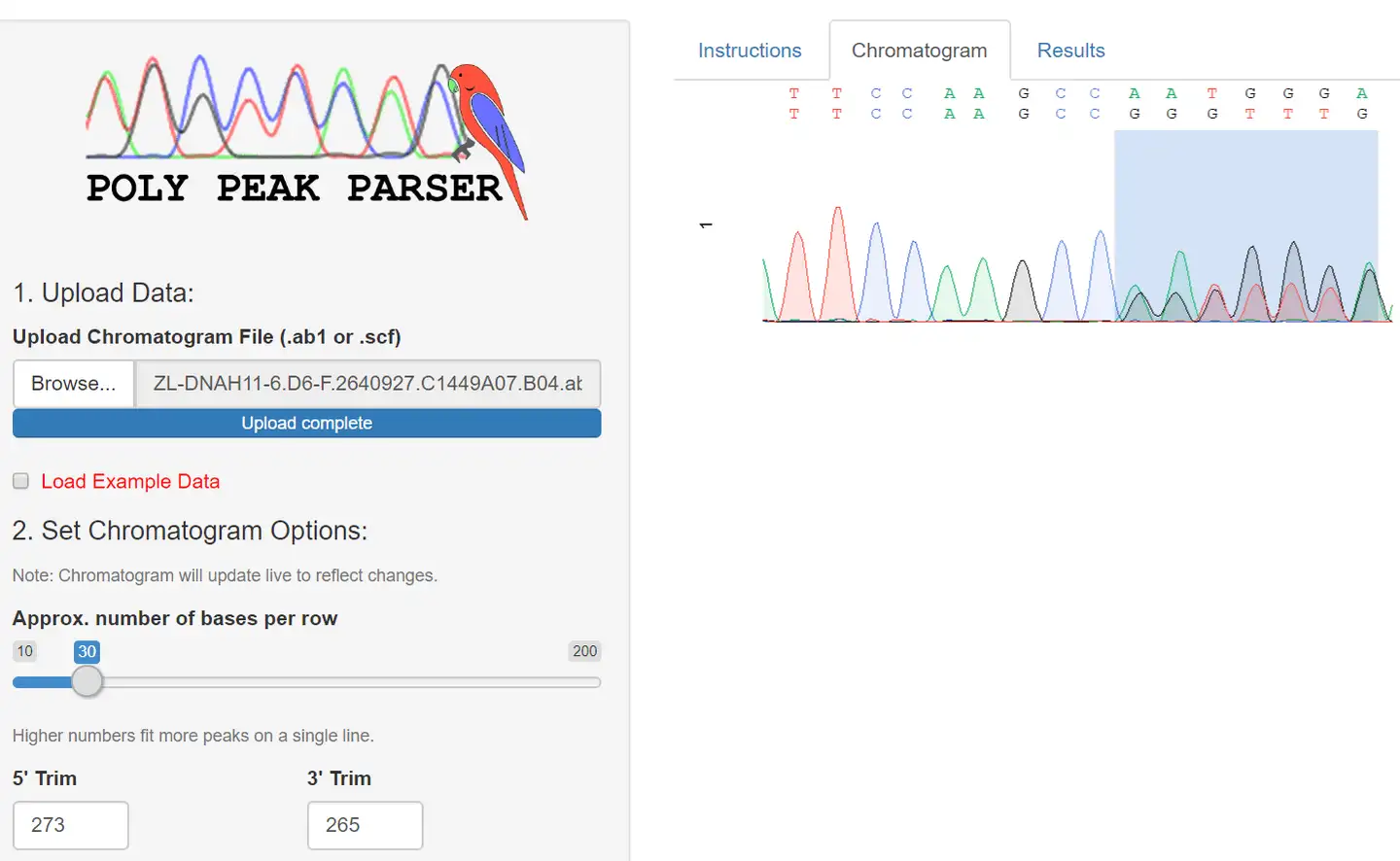
Poly peak parser: Method and software for identification of unknown indels using sanger sequencing of polymerase chain reaction products - Hill - 2014 - Developmental Dynamics - Wiley Online Library

MultiEditR: The first tool for the detection and quantification of RNA editing from Sanger sequencing demonstrates comparable fidelity to RNA-seq: Molecular Therapy - Nucleic Acids
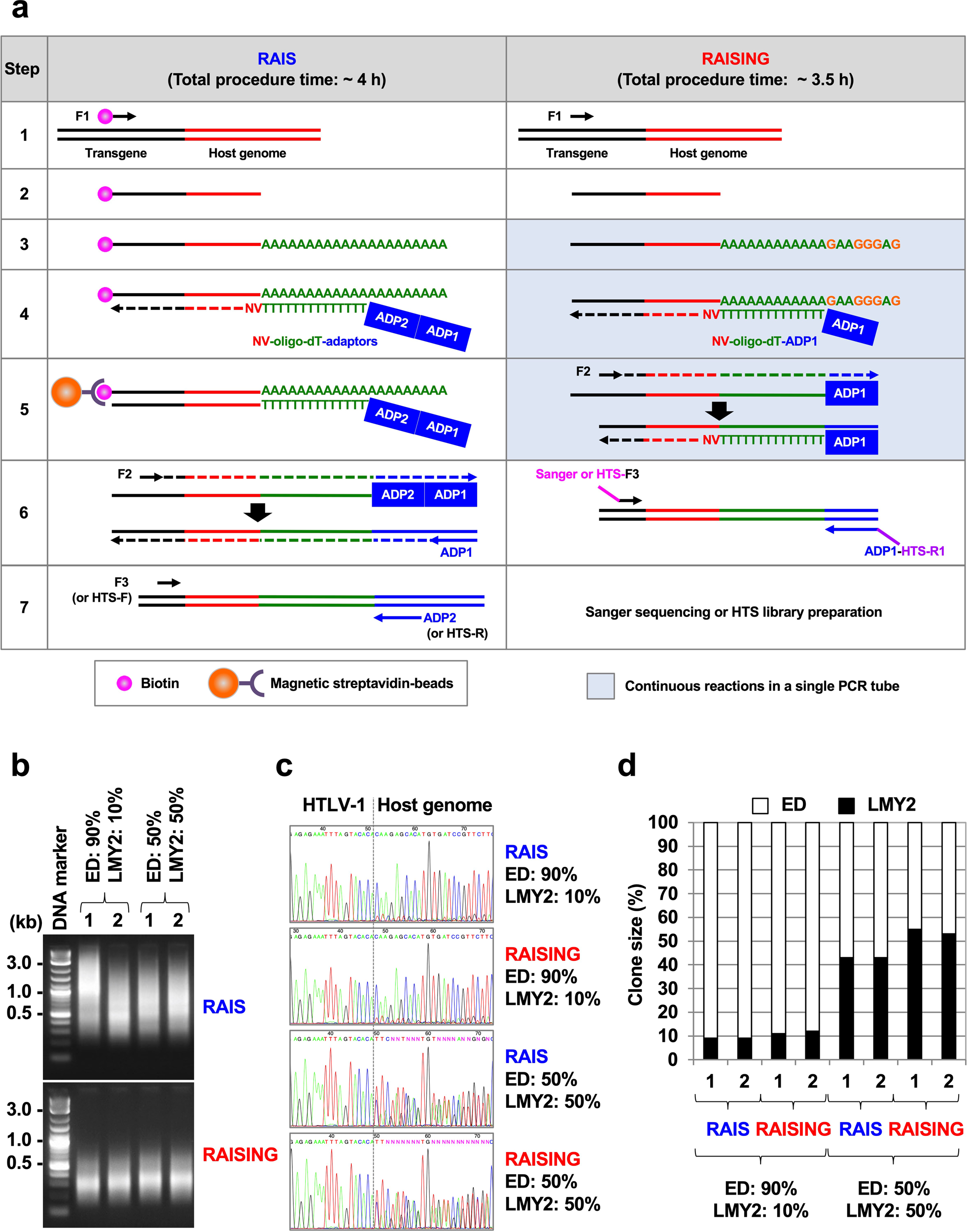
RAISING is a high-performance method for identifying random transgene integration sites | Communications Biology

MultiEditR: The first tool for the detection and quantification of RNA editing from Sanger sequencing demonstrates comparable fidelity to RNA-seq: Molecular Therapy - Nucleic Acids
mRNA processing in mutant zebrafish lines generated by chemical and CRISPR-mediated mutagenesis produces unexpected transcripts that escape nonsense-mediated decay | PLOS Genetics

MultiEditR: The first tool for the detection and quantification of RNA editing from Sanger sequencing demonstrates comparable fidelity to RNA-seq: Molecular Therapy - Nucleic Acids

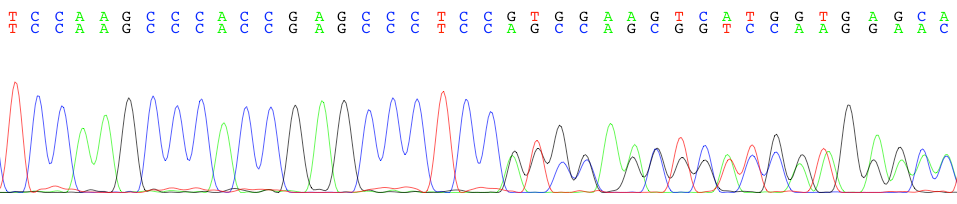


![A Python script to merge Sanger sequences [PeerJ] A Python script to merge Sanger sequences [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/11354/1/fig-3-2x.jpg)